 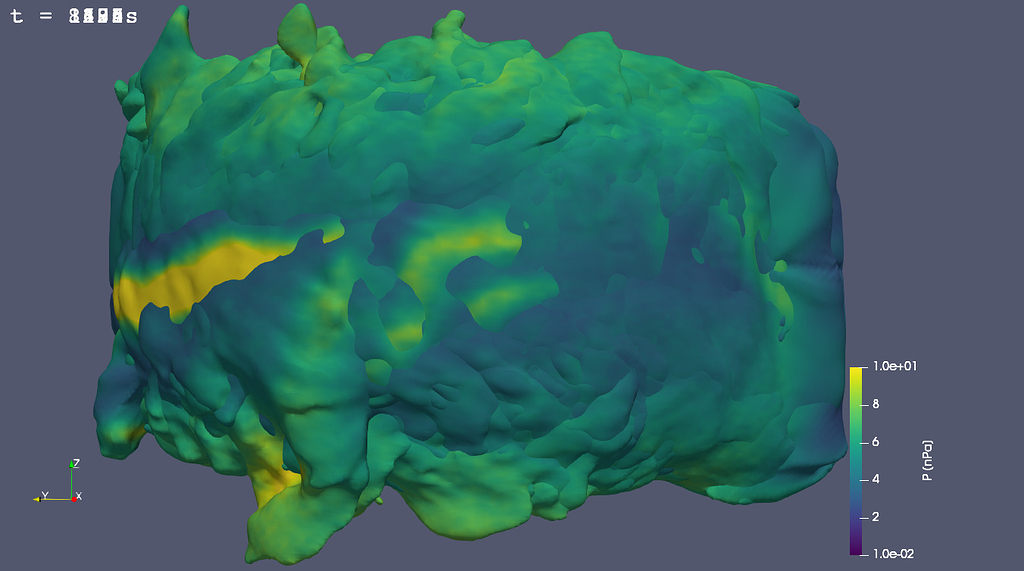
Repeat the above steps with a new instance of the. In this 9th tutorial for Paraview, I will teach you:- How to use the 'Find Data' tool to select and filter data by value- How to use the various selection op.Slice the surface data at the distance along the plane you wish to analyse.Use the ‘Calculator’ filter to calculate Cp using the formula: Cp = (p - p(freestream static))/(0.5*pho(freestream)*velocity(freestream)^2).Use the ‘Extract Surfaces’ filter to create surface data from the case file. In prost-processors such as CFD-Post from ANSYS or STAR-CCM+ from Siemens, the contour refers to a 2D plots on a specified plane or boundary surfaces.case file and select only the wing surfaces for the Upper surface in the mesh regions pannel. ParaView has many filters so looking at the filter list and deciding the logical order in which to place them can be daunting so I shall list the filters and order they were used in, to first generate the data required. I will start from the point where the results of the wing section or wing have been downloaded. Here as I was saying, you can see some nan-values (“red” in both plots) produced at the edges creating some strange artifacts not present in the numpy array.I recently wanted to view the pressure distribution of a wing section and view it as a Cp vs X/c graph and remembered that this was asked about in the forums recently so I thought I would document my workflow for future references. Here’s the state: proj-filter-testing.pvsm (608.2 KB) This is seems wierd and pretty unexpected. Then changing OutputDataType from vtkImageData to Same as input sometimes I’m getting the correct result (although with the boundary issues) but sometimes that isn’t working as well. Util.SetOutputWholeExtent(self,0,resolution,0,0])Īlso on changinng normals variable and hitting Apply makes the output on Paraview produce plots with garbage values. But if you insist on doing it using Paraview, you can. Util.SetOutputWholeExtent(self,0,0,0,resolution]) As TylerOlsen suggests, it is probably a good idea to do this particular plot using another tool. Util.SetOutputWholeExtent(self,0,resolution]) # This part of the code is to be placed in the "RequestInformation Script" entry: Ido.SetScalarComponentFromDouble(i,j,0, 0, proj_img) Ido.SetScalarComponentFromDouble(i,0,j, 0, proj_img) Ido.SetScalarComponentFromDouble(0,i,j, 0, proj_img) Weight_img = np.zeros((resolution,resolution),dtype=np.float64) 2d static analysis in workbench with apdl meshing 18 2, ansys tutorial. Proj_img = np.zeros((resolution,resolution),dtype=np.float64) The output file output.vtk is written in the same location as the input file. #All cells are assumed to be of uniform size in data Weight = np.reshape(weight,(dimensions,dimensions,dimensions),order='F') Weight = pdi.PointData #np.ones(numPts, dtype=np.float64) Ivals = np.reshape(ivals,(dimensions,dimensions,dimensions),order='F') #choose an input point data array the projection of which is to be made Ido.SetExtent(0,resolution,0,resolution,0,0) Ido.SetDimensions(resolution,resolution,1) Ido.SetExtent(0,resolution,0,0,0,resolution) Similarly, matplotlib, Seaborn, and several other. 
Ido.SetDimensions(resolution,1,resolution) in this chapter and thus sidestepping the need to worry about rendering plots with massive dataset sizes. Ido.SetExtent(0,0,0,resolution,0,resolution) Ido.SetDimensions(1,resolution,resolution) Resolution = (extent-extent,extent-extent) #pixel for the projection plot Xmin, xmax, ymin, ymax, zmin, zmax = pdi.GetBounds() of widgets for 3D interaction, and extensive 2D plotting capability. from _interface import dataset_adapter as dsaįrom _interface import algorithms as algs Vtk) create the outline of the data as polygonal mesh and show it Mesh (ugrid). 
Can you please have a look at this and also comment on the efficiency on large datasets run on parallel client-server interface? I get the correct expected image in matplotlib figure but not for the vtkImageData on Paraview (very strange behavior at the boundary). Sample At Cell Boundaries gives the most accurate plots by creating probing points at. For comparing, I was also creating a matplotlib figure on the array which I’m creating for the vtk data. The Plot Over Line filter samples the dataset attributes of the. I tried to make a vtkImageData of the filter. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
